Bootstrapper
A plugin to quickly generate dense ground truth with sparse labels
A napari plugin to quickly generate dense 3D segmentations from sparse 2D labels.
Sparse 2D annotations made in ~10 minutes on a single section can produce dense 3D segmentations that are good starting points for refinement. Based on the 2D→3D method described in Sparse Annotation is Sufficient for Bootstrapping Dense Segmentation.
For larger volumes, dedicated 3D models, and block-wise processing, see the Bootstrapper CLI.
| Dataset | Data Type | Video |
|---|---|---|
| CREMI C | 3D volumetric stack | 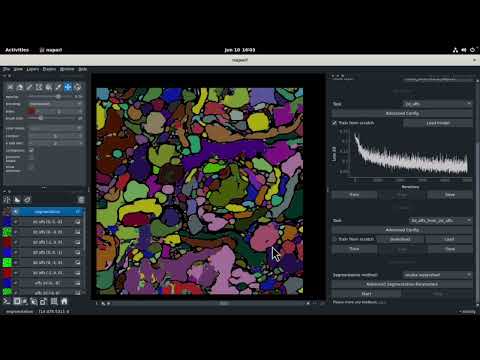 |
| Fluo-C2DL-Huh7 | 2D + time series | 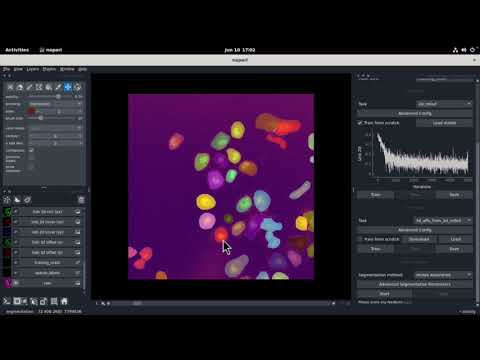 |
Getting Started
Install
conda create -n napari-bootstrapper -c conda-forge python==3.11 napari pyqt
conda activate napari-bootstrapper
pip install napari-bootstrapper
Or install the latest development version:
pip install git+https://github.com/ucsdmanorlab/napari-bootstrapper.git
Launch
conda activate napari-bootstrapper
napari
Open the Bootstrapper widget from Plugins → napari-bootstrapper.
How It Works
The plugin has four sections that follow a simple workflow:
1. Data
Load a 3D image (or 4D with a channels dimension) and create sparse 2D labels on one or a few slices. Click "Make mask" to generate a binary training mask.
We recommend using a foundation model to make sparse 2D labels, like micro-sam.
2. Train a 2D Model
Train a 2D model on your sparse labels. Three task types are available:
2d_affs— affinities2d_lsd— local shape descriptors2d_mtlsd— multi-task (both)
Or load a pretrained checkpoint.
3. Load a 3D Model
The 3D model lifts 2D predictions into 3D affinities. Pretrained weights are recommended — just click "Download".
4. Segment
Click "Start" to run the full 2D→3D inference pipeline. The output is an instance segmentation produced via mutex watershed or connected components.
Proofreading
We provide a separate widget for refining segmentations quickly. Select labels by placing points on them or entering label IDs manually. Operations can be applied per-slice (2D) or on the full volume (3D).
- Morphology — Dilate, erode, open, close, fill holes. Stenciled (3×3×3) or spherical (variable radius). Uses fastmorph.
- Merge / Split — Merge labels, split with watershed markers, or delete. Uses fastremap.
- Filter — Remove by size (min/max voxels), keep K largest, remove outliers by sigma, filter by Z-slice count, relabel connected components. Uses cc3d.
Citation
If you find this useful, please cite our preprint:
@article {Thiyagarajan2024.06.14.599135,
author = {Thiyagarajan, Vijay Venu and Sheridan, Arlo and Harris, Kristen M. and Manor, Uri},
title = {Sparse Annotation is Sufficient for Bootstrapping Dense Segmentation},
year = {2024},
doi = {10.1101/2024.06.14.599135},
URL = {https://www.biorxiv.org/content/10.1101/2024.06.14.599135v2},
}
Issues
If you encounter any problems, please file an issue along with a detailed description.
Acknowledgements
- micro-sam — Making foundation models like SAM accessible to the community.
- empanada-napari — Proofreading widget inspiration.
- napari-cellulus — Development scaffolding.
Funding
Chan-Zuckerberg Imaging Scientist Award DOI https://doi.org/10.37921/694870itnyzk from the Chan Zuckerberg Initiative DAF, an advised fund of Silicon Valley Community Foundation (funder DOI 10.13039/100014989).
NSF NeuroNex Technology Hub Award (1707356), NSF NeuroNex2 Award (2014862)
Version:
- 0.3.0
Last updated:
- 2026-03-02
First released:
- 2025-05-23
License:
- Copyright (c) 2025, Vijay Venu...
Supported data:
- Information not submitted
Plugin type:
Open extension:
Save extension:
Python versions supported:
Operating system:
- Information not submitted
Requirements:
- numpy
- scipy
- scikit-image
- torch
- zarr<3
- numba
- gunpowder
- magicgui
- qtpy
- pyqtgraph
- matplotlib
- napari
- tqdm
- lsds
- mwatershed
- fastmorph[spherical]
- connected-components-3d
- fastremap
- tox; extra == "testing"
- pytest; extra == "testing"
- pytest-cov; extra == "testing"
- pytest-qt; extra == "testing"
- napari; extra == "testing"
- pyqt5; extra == "testing"




